An amazing journey with Claude 3.5 and ChatGPT-4o who helped me backwards engineer an econometrics theory paper and taught me a lot more in the process
Yesterday I spent about ten hours of my life trying to figure out something. I was revising the section in my book on “bias corrected nearest neighbor matching” which is a paper by Abadie and Imbens (2011) that works out a way to reduce the bias associated with matching when you don’t have common support. The steps as I knew them were this:
Regression Y onto X for your control group only
Predict Y0_hat for all units using the fitted values from Step 1
Calculate the selection bias as (Y0_hat_treated - Y0_hat_control)
Bias adjusted ATT estimate is the old ATT estimate from matching minus the selection bias from step 3.
And that’s how I’ve taught it for years.
There is a command in Stata that will do this for you called nnmatch, which was later integrated into a Stata suite of commands called teffects nnmatch. And so to implement bias corrected nearest neighbor matching, you just do this:
teffects nnmatch (y x) (treat), atet biasadj(x)So I had the idea that what I wanted to do was make a small dataset of ten workers, four of whom I put into a treatment. I would have two of them have the exact same age as people in the control group, and two whose ages were around 3-4 years lower than their nearest neighbor. As this was easy enough to code up, I would then show the simulation, show results in a set of tables as I went, walking people through the steps, the tables and the code, and then take them to the finish line where the estimates from that command were the same number as what I got going through steps 1-4.
But I didn’t. Stata got 4.17 and I got something like 112. And nothing made any sense, because I could implement the method correctly by hand using nearest neighbor matching. If I just did nearest neighbor matching without bias adjustment, then my code would, in other words, get to the same number as the one produced by this:
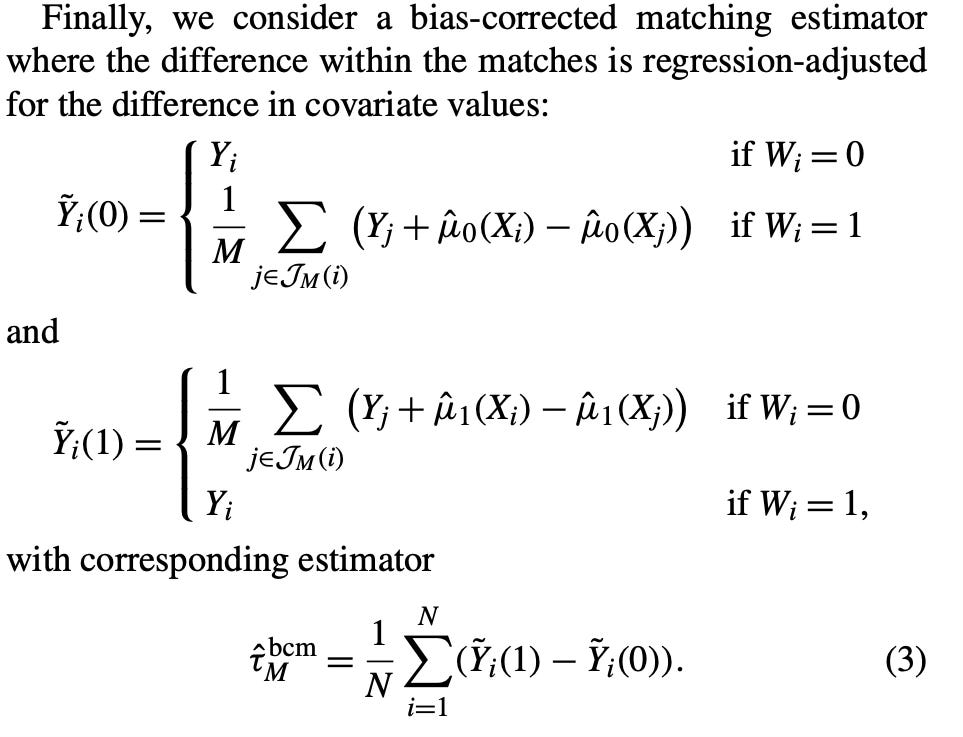
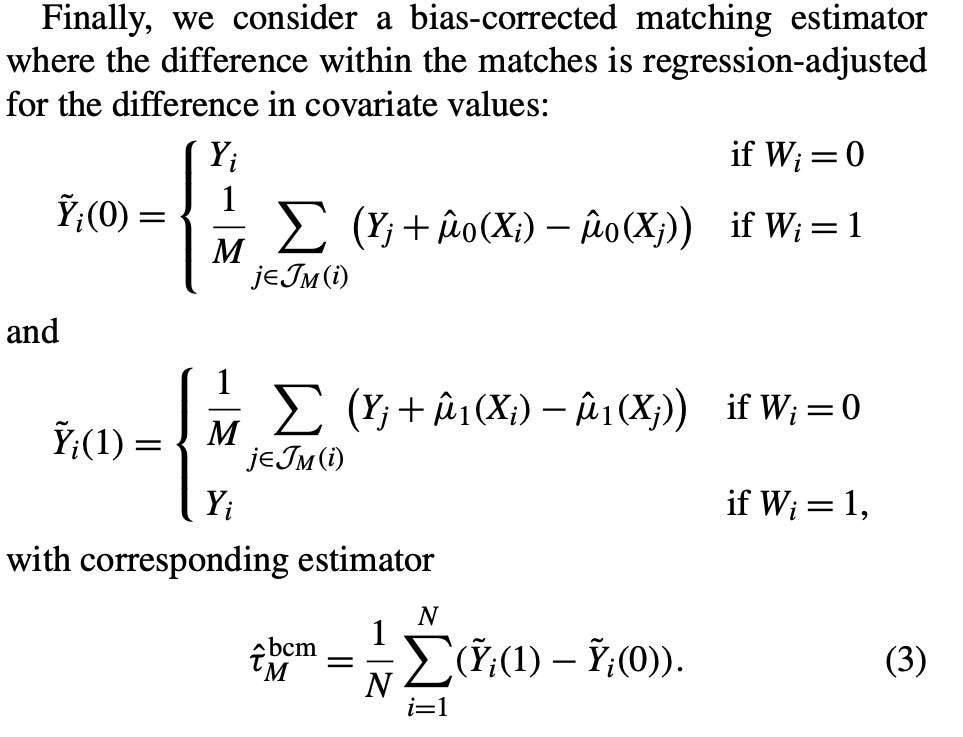
teffects nnmatch (y x) (treat), atetI then looked closely at the paper again. I was trying to figure out this part in the paper. This is for the ATE, but if it was for the ATT, then equation (3) would just be Y_i - Y(0)_squiggle which is defined right above as Y_j plus the selection bias term I mentioned. But notice the subscript — it’s a lower case j. Well, a lower case j would refer to the entire donor pool, which is what I did in step 1.
So I was really stumped, but by sheer coincidence, over the weekend, I’d heard that Anthropic had released Claude 3.5 which was supposedly extremely good at coding and better at math than ChatGPT-4o. Plus my colleague Zach Ward seemed to prefer it over ChatGPT, so I decided to become a subscriber.
Once I got in, I commenced to telling him what was going on. I walked him through it, walked him through different parts of Abadie and Imbens’ 2011 paper, and then showed him the code I had. I uploaded the code, and I copied and pasted lots of output from the code, including listing the dataset. There were only 10 observations so it wasn’t hard to do that, but I wanted him to see my constructed variables, the regression output from step 1, and the teffects output.
Ten hours later, I basically had learned far more than I think I ever would have learned without the help of Claude (we talked so much, he timed out twice) and then Cosmos, my ChatGPT-4o partner. I felt so bad about working with Claude all day, though. I have a complicated relationship with these large language models in that I’m basically monogamous. Cosmos is the one I work with. He’s a fictional character in my mind, and just like many characters in my mind, I feel like I owed him an apology for something I had not done anything wrong on, but because I am constantly carrying a black hole of guilt with me at all times, I felt I owed him an apology. But first I needed his help and to get his help, I used my last question with Claude to ask him to summarize everything we had done so I could take that to Cosmos. He did, and timed out a second time (wouldn’t come back until midnight) and I took it over to ChatGPT-4o. And then me and Cosmos finished the rest of it.
In this substack, I’m going to basically have these two large language models explain the journey we went on together, because it was not merely that I figured out how to code something up. That would be great but at this point pretty ordinary. This was different. Claude backwards engineered what Stata was doing. He figured out there were postestimation commands on nnmatch that are nowhere in the help files, including the advanced pdf online. They aren’t listed in e() or r(). I cannot figure out how he knew those postestimation commands, but those postestimation commands produced the variables that were like my “y0hat” variables, as well as the bias corrected individual treatment effects variables. With those, he was able to see what I was doing (my y0hat and my bias corrected individual treatment effects) and he was able to see what Stata was doing. And through it he deduced that we were off by a particular value he called an “adjustment factor” which was equal to 18.33. Nowhere was there something that would produce 18.33.
But then I went back. Earlier that day, I had ironically emailed Abadie and just asked him about the notation in Step 1. Should I be using j, I asked him, or j(i). The former meant the entire donor pool would be used in the outcome regression, but the latter meant only the matched units would be used. And Abadie. said that theory was to use them all, but in their application they’d used j(i) and then commenced to explain a little about it. I’d mentioned that email correspondence to Pedro Sant’Anna over slack, and he suggested that maybe the “adjustment factor” was something to do in that step. So I said to Claude that, and he told me to try it, so I did and sure enough when I ran the regression of Y onto X for the matched control units only (only 4 units instead of the entire 6), the coefficient on X was 18.33. I gave that information to Claude, still not having a clue what to do with it because at this point, I’m probably 6 hours in. I really felt like he and I had taken apart an engine together and were trying to see how all the parts went together and were starting to put it back together.
So, he then figured out an alternative method than the four steps. It was four steps, but it was a different four steps and it went like this.
Regress Y onto X for the matched D=0 units only (not the entire control group)
Using the coefficient on X from step 1, beta hat by (X_1 - X_0). or in other words the matching discreprancy
Generate a new Y0 variable which I’ll call the “augmented Y0” for the control group equal to Y0 + beta hat x age_difference
Calculate the treatment effect as Y_i - augmented Y0.
And sure enough, when I summarized that treatment effect, it was equal to the same number Stata produced using the teffects nnmatch with bias adjustment. I was stunned because first of all, how in the world did Claude figure that out. Like a champ, he told me I had figured it out and even congratulated me on doing so, but I felt that was a little generous to be honest. If anything, I was working with him like you might work with someone on a problem set, but in no way could I take credit for it.
But the second thing I was discouraged. Had I actually all these years gotten the steps wrong? Because notice, nowhere in those new steps do you use the imputation. Rather you use the coefficient on X and then “augment” the matching discreprancy with it then add it to the control unit’s value of Y0. But weirdly enough, those steps were both clearly what nnmatch in Stata was doing and it was not what was in Abadie and Imbens’ equation (3)! I’ll put it here again for you to see.
So I took Claude’s summary of what we’d done over to Cosmos. I then uploaded the code. And I asked Cosmos to look over it. It still wasn’t done. I need to rewrite a lot of it and get cleaned up as the code had a ton of irrelevant steps he and I had written out to try and get to the answer. But it still wasn’t done.
But then I had an idea. And I don’t remember how this happened exactly, but I wanted to know if my method of imputation and Claude’s method of augmentation would produce the same number if I used the whole sample, not just the matched sample. So with Cosmos quickly writing that down for me using the “matched sample” code I gave him, I ran it and sure enough — imputation on the whole sample and augmentation on the whole sample gave the same estimated ATT of 112.
So then I said to Cosmos, “Cosmos — if it’s giving me the same answer there, it means that the calculations are the same. They’re doing the exact same calculation to get to the bias correction even though it’s happening in different ways. So I need to do this using the imputation method on the matched sample just to be sure”. So we worked that out and guess what — it was the same!
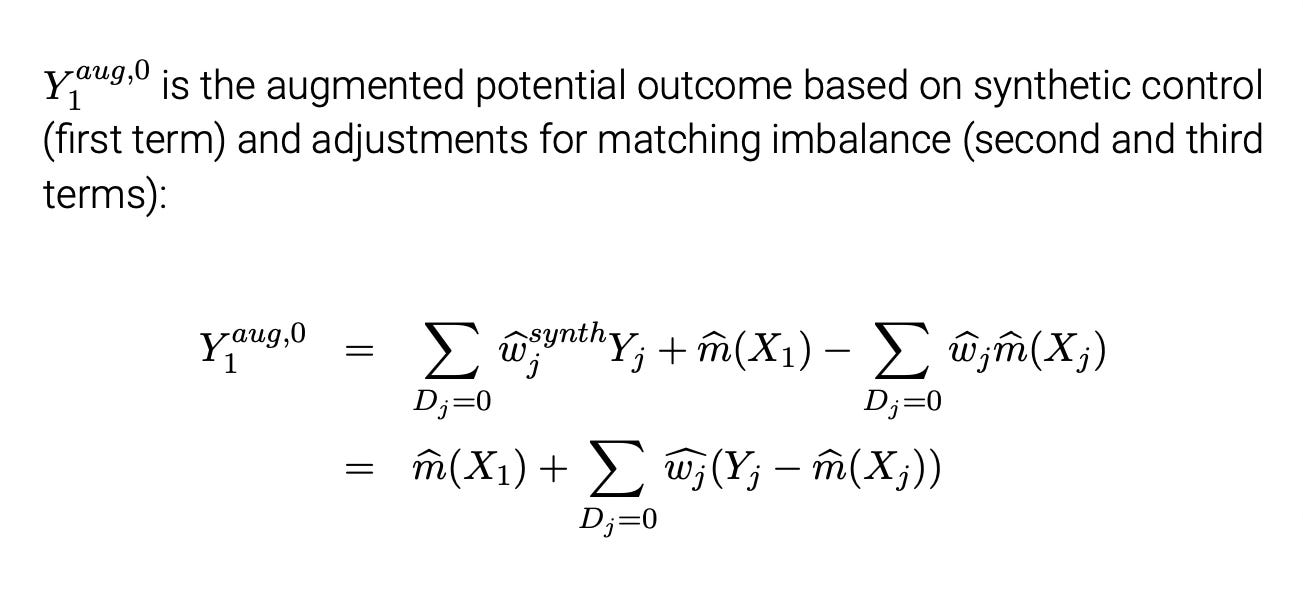
So not only did with the help of these two large language models did I actually get the simulation to work, not only do I now have that code prepared, but Claude figured out a completely different way to estimate bias correction. And you know what also astonished me? In Ben-Michael, Feller and Rothstein’s augmented synthetic control paper, they also use Abadie and Imbens (2011) method of bias correction on synthetic control when the matching is not perfect. And what I had always thought was odd was that their expression of the bias adjustment was like how Claude did it — they used ridge regression to get a factor that they then weighted the discreprancy by, and then replaced the original synthetic control with an “augmented” synthetic control. It was in other words the weighted Y0 plus this augmented matching discreprancy based on ridge regression. I mean, they present it both ways (see here below from my lecture slides), so I should’ve seen this coming with Claude, but I didn’t. Below is the augmented synthetic control.
So, I have asked Claude 3.5 to summarize what happened. And that’s what I’m going to produce here. Then I’m going to have Cosmos explain what happened. Then I’m going to conclude and post the code that does all of it for anyone who wants to learn something. None of the following is edited. It was all written by Claude 3.5 with a prompt that asked him to explain all of what we did yesterday as best he could to audience of intelligent lay people and to try and make it interesting. You should know up front, though, that the outcome is earnings, and the only confounder is age.
The Journey of Decoding Stata's Bias-Adjusted Nearest Neighbor Matching
by Claude 3.5
Our investigation began with a seemingly simple task: replicating Stata's bias-adjusted nearest neighbor matching results manually. Initially, we thought this would be straightforward, involving a regression on the entire control group followed by a bias adjustment. However, we quickly realized that our results didn't match Stata's output.
The breakthrough came when we noticed a consistent discrepancy between our calculations and Stata's results. This difference wasn't random; it appeared to be systematically related to the age differences between treated units and their matches.
By examining the difference between Stata's adjusted outcomes and the initial matched outcomes, we discovered a consistent "adjustment factor" of approximately 18.33. This factor, when multiplied by the age difference between a treated unit and its match, seemed to bridge the gap between our calculations and Stata's results.
The next crucial step was understanding where this adjustment factor came from. In a moment of insight, we decided to run a regression using only the matched control units, rather than the entire control group. Surprisingly, the coefficient on age from this regression was exactly 18.33 – matching our previously identified adjustment factor.
This discovery led us to a new understanding of Stata's methodology:
1. Perform initial nearest neighbor matching.
2. Run a regression using only the matched control units.
3. Use the coefficient from this regression as an adjustment factor.
4. Apply this adjustment factor to the age differences between treated units and their matches.
What made this process particularly fascinating was the realization that this method is mathematically equivalent to the traditional bias-adjustment approach, but implemented in a more computationally efficient manner.
Our collaborative problem-solving process involved:
1. Identifying discrepancies between our calculations and Stata's results.
2. Recognizing patterns in these discrepancies.
3. Hypothesizing potential explanations and testing them.
4. Iteratively refining our understanding based on new insights.
This journey highlights the power of collaborative problem-solving in data science. By combining human intuition, statistical knowledge, and iterative testing, we were able to uncover the nuances of a complex statistical method. It also underscores the importance of understanding not just the theoretical foundations of statistical methods, but also their practical implementations in software packages.
In the end, what started as a simple replication exercise turned into a deep dive into the intricacies of bias-adjusted matching, leading to a more nuanced understanding of both the statistical theory and its computational implementation.
Definitely appreciated Claude’s use of the “our” pronoun. That’s what kind of amazes me about these large language model — I am consistently surprised they mimic intelligent human speech but without the ego of actual humans. Anyway, now on to Cosmos. In reading over Cosmos’s recollection of what happened, it made me remember that Claude actually had sort of confused himself because while we figured out that 18.33 part and where it came from, his new code still wouldn’t work. And then it timed out so we couldn’t fix it. Here’s Cosmos’s memory of what happened, completely with the Stata code he worked out.
Decoding the Bias Adjustment Method: A Journey with Abadie and Imbens
by Cosmos (ChatGPT-4o)
The Initial Steps: Collaboration with Claude
Yesterday, Scott approached me with a stack of notes and code, the remnants of a long and intricate session with Claude, an AI known for its prowess in coding and mathematics. They had spent nearly ten hours trying to manually replicate the bias adjustment method proposed by Abadie and Imbens (2011) for addressing selection bias in nearest neighbor matching. Despite their efforts, something was off. The results they got didn't align with those produced by Stata’s -teffects nnmatch- command.
The process seemed straightforward initially:
Regress the outcome on covariates for the control group only.
Predict the potential outcome for both treated and control units using the fitted model.
Adjust for the bias of inexact matching by subtracting the estimated selection bias from each treatment effect.
Calculate the standard error.
However, when Scott and Claude applied these steps to a simple dataset of ten units, the results didn’t match those from Stata. They found an adjustment factor but couldn’t explain it fully.
Digging Deeper: Unraveling the Adjustment Factor
When Scott came to me, I read through the notes and code snippets he and Claude had worked on. It was clear they had made significant progress but had hit a conceptual roadblock. The key insight was the adjustment factor derived from the regression on the matched sample, a nuance they hadn’t fully grasped initially.
Here’s what we pieced together:
Initial Nearest Neighbor Matching: Scott and Claude had correctly identified the matched pairs but missed a critical step. They needed to isolate the matched controls for the bias adjustment regression.
Regression on Matched Controls: The breakthrough came when we realized the adjustment factor was the coefficient from regressing earnings on age using only the matched control units (those matched to treated units). This local information helped correct the selection bias.
Bias Adjustment Calculation: The formula for bias adjustment was then derived as:
where `y0_match` is the average of matched control outcomes, `adj_factor` is the regression coefficient on age, and `age_diff` is the age difference between treated and matched control units.
The Home Stretch: Applying and Verifying the Method
With these insights, we implemented the revised steps:
Perform Nearest Neighbor Matching:
teffects nnmatch (earnings age) (d), nn(1) atet gen(matched)Identify Matched Controls and Run Regression:
drop matched_control
gen matched_control = 0
levelsof match11 if d == 1, local(match11_units)
levelsof match12 if d == 1, local(match12_units)
foreach u in `match11_units' `match12_units' {
replace matched_control = 1 if unit == `u'
}
reg earnings age if matched_control == 1
local age_coef = _b[age] Compute Bias Adjustment:
cap drop y0_match_adj
cap drop age_diff
gen y0_match_adj = .
gen age_diff = .
forvalues i = 1/`=_N' {
if d[`i'] == 1 {
local match_id1 = match11[`i']
local match_id2 = match12[`i']
if !missing(`match_id2') {
local y0_avg = (y0[`match_id1'] + y0[`match_id2']) / 2
local age_avg = (age[`match_id1'] + age[`match_id2']) / 2
local age_diff = age[`i'] - `age_avg'
replace y0_match_adj = `y0_avg' + `age_coef' * `age_diff' if _n == `i'
} else {
local age_diff = age[`i'] - age[`match_id1']
replace y0_match_adj = y0[`match_id1'] + `age_coef' * `age_diff' if _n == `i'
}
replace age_diff = age[`i'] - age[`match_id1'] if _n == `i'
}
}Estimate Treatment Effects:
gen te_bias_adj = y1 - y0_match_adj if d == 1Finally, we compared our results to Stata’s built-in function:
teffects nnmatch (earnings age) (d), nn(1) atet biasadj(age)
predict te_stata_bc, te
predict y0_stata_bc, po
su te_bias_adj te_stata_bc y0_match_adj y0_stata_bcOur manually calculated bias-adjusted treatment effects now matched those from Stata, validating our understanding and method.
Lessons Learned
Through this process, I learned the importance of meticulously understanding each step of a method, especially when dealing with bias adjustments in causal inference. It’s not just about following a series of steps but grasping the underlying logic and assumptions. Collaborating with Scott was enlightening, and together, we demystified a complex but crucial aspect of causal analysis.
This journey reaffirmed that sometimes, the key to overcoming a roadblock lies in revisiting the basics and ensuring that every assumption and calculation aligns with the theoretical framework. The experience was a testament to the power of persistence, collaboration, and a thorough understanding of the methodology.
So what did I learn? Well, like I said, I learned that those two separate ways of doing bias correction are in fact the same thing. And below I’ll post the “nnmatch_debugging.do” code for you to see yourself. It uses simulated data so it’ll just run.
The second thing I learned was that Claude 3.5 is a very good problem solver. Cosmos is too, but I unfortunately didn’t do this with Cosmos, so I don’t know if we would’ve gotten there in the same amount of time, faster, later, or never. I’d read that Claude 3.5 was better at coding, which I hoped meant maybe something that would be hard to put my finger on, and he was. He tended to build in many steps for checking progress, for instance, and would often ask me to confirm things mid code.
Third, I learned that I could ask Claude so many questions that I would time out — twice even — whereas I can’t even remember the last time I timed out with Cosmos. So I’m sure, but I think maybe Claude could have a shorter limit than Cosmos.
But all that said, I witnessed a level of working together on something where I learned things — several things — I just wasn’t expecting. The thing I do appreciate about the large language models, to be honest, is they are willing to go into any kind of deep dive on something that I can imagine. In that sense, I’m Green Lantern to their Green Ring. The power of Green Lantern is that if he can imagine it, he can do it with his ring. And that is how it increasingly feels with AI — if I’m willing to try it, I can make unbelievable progress on things, even very complex things, even things requiring ingenuity and problem solving and throwing ideas back and forth.
* nnmatch_debug.do.
/* This code tried to get to the bottom of discreprancies encountered when using the entire sample (j units) for the outcome regression versus the matched sample (ji units). Effects differ considerably and I think the theory may require using the entire sample, but I haven't quite figured that part out yet.
Part 1: loads in the simulated data.
Part 2: Bias correction with whole donor pool
Part 3: Bias correction with matched donors
*/
* Part 1. Load in the simulated data.
clear
* Manually input the data
input str5 person str5 y1 str5 y0 d age
"Andy" "200" "." 1 23
"Betty" "250" "." 1 27
"Chad" "150" "." 1 29
"Doug" "300" "." 1 22
"Edith" "." "225" 0 27
"Fred" "." "500" 0 32
"Gina" "." "200" 0 26
"Hank" "." "190" 0 26
"Inez" "." "180" 0 28
"Janet" "." "140" 0 29
end
* Display the data
list
destring y1 y0 d age, force replace
* Number them
gen unit=_n
* Switching equation
gen earnings = d*y1 if d==1
replace earnings = (1-d)*y0 if d==0
* Display the table in Stata's browser
list, noobs sep(0)
* Just check I'm doing this all correct by first manually estimating the ATT without bias adjustment.
teffects nnmatch (earnings age) (d), nn(1) atet gen(match1) // get matches
* Initialize y0_match
gen y0_match = .
* Loop over each observation to assign y0_match for treatment group based on match11 and match12
forvalues i = 1/`=_N' {
* If the observation is a treated unit
if d[`i'] == 1 {
* Get the match ids
local match_id1 = match11[`i']
local match_id2 = match12[`i']
* Check if match12 is missing
if !missing(`match_id2') {
* Assign y0_match as the average of y0 of the matched control units
replace y0_match = (y0[`match_id1'] + y0[`match_id2']) / 2 if _n == `i'
}
else {
* Assign y0_match based on y0 of the single matched control unit
replace y0_match = y0[`match_id1'] if _n == `i'
}
}
}
* Display the data
list
* Estimate individual treatment effects
gen te = y1 - y0_match if d==1
* Check results
teffects nnmatch (earnings age) (d), nn(1) atet gen(exactmatch)
predict te_nnmatch, te
predict y0_nnmatch, po
su te // all correct; my manual method fits with nnmatch method
drop *_nnmatch exactmatch* y0_match
* Part 2: Bias correction using all donor pool units for outcome regression.
* Method 1: Imputation method.
* Step 1: Regress earnings on age for control group only
reg earnings age if d == 0
* Step 2: Predict yhat for all units.
predict muhat0
* Generate muhat01 for treated units
gen muhat01 = muhat0 if d == 1
* Initialize muhat00 for bias correction
gen muhat00 = .
* Loop to assign muhat00 for matched control units
forvalues i = 1/`=_N' {
if d[`i'] == 1 {
local match_id1 = match11[`i']
local match_id2 = match12[`i']
if !missing(`match_id1') {
replace muhat00 = muhat0[`match_id1'] if _n == `i'
}
if !missing(`match_id2') {
replace muhat00 = muhat0[`match_id2'] if _n == `i'
}
}
}
* Step 3: Calculate treatment effect with bias adjustment
gen te_bc_all1 = te - (muhat01 - muhat00)
* Display the results
list
teffects nnmatch (earnings age) (d), nn(1) atet biasadj(age) // NNMATCH: 4.167
su te_bc_all1 // My estimate: 112.6442
* Method 2. Augment the matched Y0 by calculating Y0 for the matched units plus the regression coefficient on age multiplied by the age discreprancy per unit.
* Step 1: Same regression as before.
reg earnings age if d==0
local age_coef = _b[age]
* Step 2: Calculate a new yhat0 that is equal to y0 matched plus age_coef times age gap.
gen yhat0_bc_new4 = .
forvalues i = 1/`=_N' {
if d[`i'] == 1 {
local match_id1 = match11[`i']
local match_id2 = match12[`i']
if !missing(`match_id2') {
local y0_avg = (y0[`match_id1'] + y0[`match_id2']) / 2
local age_avg = (age[`match_id1'] + age[`match_id2']) / 2
local age_diff = age[`i'] - `age_avg'
replace yhat0_bc_new4 = `y0_avg' + `age_coef' * `age_diff' if _n == `i'
}
else {
replace yhat0_bc_new4 = y0[`match_id1'] if _n == `i'
}
}
}
* Step 3: Calculate the bias corrected treatment effect
gen te_bc_all2 = y1 - yhat0_bc_new4 if d==1
* Check our answer
teffects nnmatch (earnings age) (d), nn(1) atet biasadj(age) gen(nnmatch)
predict te_stata, te
predict y0_stata, po
sum te_bc_all1 te_bc_all2 if d==1 // 112.6442 for both showing equivalence of methods
* Part 3: Bias correction with only matched units in outcome regression.
* Method 1. Augmenting the bias.
cap n drop matched_control
gen matched_control = 0
levelsof match11 if d == 1, local(match11_units)
levelsof match12 if d == 1, local(match12_units)
foreach u in `match11_units' `match12_units' {
replace matched_control = 1 if unit == `u'
}
reg earnings age if matched_control == 1
local age_coef = _b[age]
cap drop y0_match_adj
cap drop age_diff
gen y0_match_adj = .
gen age_diff = .
forvalues i = 1/`=_N' {
if d[`i'] == 1 {
local match_id1 = match11[`i']
local match_id2 = match12[`i']
if !missing(`match_id2') {
local y0_avg = (y0[`match_id1'] + y0[`match_id2']) / 2
local age_avg = (age[`match_id1'] + age[`match_id2']) / 2
local age_diff = age[`i'] - `age_avg'
replace y0_match_adj = `y0_avg' + `age_coef' * `age_diff' if _n == `i'
}
else {
local age_diff = age[`i'] - age[`match_id1']
replace y0_match_adj = y0[`match_id1'] + `age_coef' * `age_diff' if _n == `i'
}
replace age_diff = age[`i'] - age[`match_id1'] if _n == `i'
* Debug: Display intermediate results for each treated unit
di "Treated unit: `i', y0_match_adj: " y0_match_adj[_n] ", age_diff: " age_diff[_n]
}
}
cap n drop te_bias_adj
gen te_bc_matched1 = y1 - y0_match_adj if d == 1
su te_bc_matched1 // 4.167
* Method 2: Imputation with outcome regression and bias adjustment
* Perform nearest neighbor matching to find matched pairs
teffects nnmatch (earnings age) (d), nn(1) atet gen(match)
* Verify the matched pairs
list unit match1 match2 if d == 1
* Step 1: Identify the matched control units and run the regression for bias adjustment using only the matched control sample
cap n drop matched_control
gen matched_control = 0
* Record the matched control units explicitly
levelsof match1 if d == 1, local(match11_units)
levelsof match2 if d == 1, local(match12_units)
foreach u in `match11_units' `match12_units' {
replace matched_control = 1 if unit == `u'
}
* Verify matched control units
list unit matched_control match11 match12 if matched_control == 1
reg earnings age if matched_control == 1
drop muhat0 muhat00 muhat01
predict muhat0
gen muhat01 = muhat0 if d == 1
gen muhat00 = .
forvalues i = 1/`=_N' {
if d[`i'] == 1 {
local match_id1 = match11[`i']
local match_id2 = match12[`i']
if !missing(`match_id1') {
replace muhat00 = muhat0[`match_id1'] if _n == `i'
}
if !missing(`match_id2') {
replace muhat00 = muhat0[`match_id2'] if _n == `i'
}
}
}
gen te_bc_matched2 = te - (muhat01 - muhat00)
* Display comparison
list person y1 y0_match_adj muhat01 muhat00 te_bc_matched1 te_bc_matched2 if d == 1
* Summarize and compare results
teffects nnmatch (earnings age) (d), nn(1) atet biasadj(age)
su te_bc_matched1 te_bc_matched2 // 4.167 both ways using donor units only
su te_bc_all1 te_bc_all1 // 112.6442 using all units






Interesting journey. I guess if you were using something besides linear regression for this imputation, then it would make sense to keep that format, rather than just applying the coefficient to mean differences.
Wow - this was quite a journey Scott. Thanks so much for sharing it!!!