If Non-Standard Errors Are Measuring Real Uncertainty, Should We Report Them?
First — thank you all for following along on this series. I’ve been writing about Claude Code since December 13th, 2025, and while today’s post is connected to that series, I decided to make it a more general post because it’s not technically about Claude Code. I mean it is and it isn’t. It’s a more general question about statistics that Claude Code made me wonder about I guess is my point and therefore I wanted to make this one a more general open ended one that doesn’t get catalogued as a Claude code post.
That said, it has genuinely been a labor of love, and the support from paying subscribers has meant a lot. And if you’re new to the substack, I wanted to say that I suspect if you subscribe, you’ll get nearly daily updates from me. Most of the time it’ll be one of four kinds of posts too:
Posts about AI, and Claude Code specifically. These are usually focused on “using AI agents for practical empirical research”. They’re not usually thought pieces though sometimes they are. But mainly I’m trying to illustrate using CC for actual practical empirical work,
Causal inference. Traditionally I write explainers on here where I’m talking about causal inference methodologies or elucidating estimators or basic tasks around them. I have a new book coming out this summer, plus I do talks on causal inference, so you’ll also hear me just talking about things related to that too.
A long list of links to articles and what not I’ve been reading that week, usually called “Closing Tabs” because it’s articles I’ve left open in my browser. They are usually links to articles about love, relationships, causal inference, pop culture, AI and then random stuff.
None of the above. This has included a lot of history of thought stuff but it’s also been books I’ve been reading, like the Courage to be Disliked, about Adlerian psychology.
So that’s more or less the deal. The Claude code posts are always free; the open tabs are free. The causal inference and the #4 are randomized paywalls on the date of the post. And then everything eventually goes behind a paywall after around 4 days. I also post podcast episodes every now and then but I’m behind on season five to that, and it’s always free.
And that’s it. That’s the gist of this substack. So if you aren’t a paying subscriber yet, these Claude Code posts are free for about four days before they go behind the paywall, so hopefully that’s enough to show you what’s going on. But if this is the day you feel like becoming a paid subscriber — at $5/mo, I think it’s a deal.
But today’s post will have to do with statistics. While I got the idea from my Claude code series, it is not about that. It’s about something I’d been thinking about for a while and am now more openly wondering about the underlying uncertainty implied by it. So it won’t go with Claude code.
Many analysts, one dataset, one treatment assignment
I want to talk about something that’s been sitting in the back of my mind since I started this series, and which I think the experiment I’ve been running the last few weeks has started to make concrete. It’s about a concept called the many-analyst design. And it’s about what I think Claude Code accidentally lets you do with it.
If you’ve been following along, you know I’ve been running a structured experiment. Same dataset. Same estimator — Callaway and Sant’Anna, not-yet-treated comparison group. Same research question. The only thing I let vary was covariate selection and software package. I gave Claude Code five packages: two in Python, two in Stata, one in R. Three trials per package. Fifteen total runs of the same study, holding almost everything fixed, and letting only one dimension of researcher discretion vary.
What you get is a forest plot showing the distribution of all the estimates from the same dataset coming from different researchers. But what is the variation? Well, it’s not the sampling distribution of the estimator as that comes from iid sampling with hypothetically constructed alternative samples. That’s one of the traditional sources of uncertainty in statistics and it isn’t that one.
It also isn’t uncertainty in the treatment assignment, which is a design based randomization dating back to Fisher 1935, and the lady tasting tea. That’s a second source of variation in treatment estimates one could construe and it isn’t that one either,
Under both iid sampling and design approaches, you are able to construct intervals and do hypothesis tests that help quantify the uncertainty in your estimates. They feed into either direct analytically derived intervals, or you can use computational resampling style procedures to get them. They mean different things, but they’re both efforts to quantify uncertainty around point estimates and have an effort to capture confidence around some sought-after answer to a target question.
It is not clear what we might be learning from a forest plot of estimates don’t by many analysts. Except there does seem to be a distribution of estimates one could construe exists, and which could therefore be undertaken with Claude code. This is only me thinking out loud, but bear with me as I do it.
Three kinds of uncertainty: sampling
There’s a standard way to think about uncertainty in empirical work, and it really only has two flavors.
The first one everyone learns in statistics and econometrics: sampling uncertainty. You drew one sample from a population. That’s your dataset. It’s a fixed size. It has specific people inside it. It would seem to you like it is the only dataset that could’ve ever existed because it is the only dataset that ever existed, but there could’ve been others. Thus there are counterfactuals in sampling based inference but the counterfactuals are the counterfactual samples based on a randomizing process of constructing samples.
The point is that it isn’t the only dataset that could’ve ever existed. As this sample contains real people, picked randomly, from a larger pool called “the population”, you could’ve had a different dataset of the same size with somewhere between a slightly different to an entirely different group of people in it. And if you had done your procedures on all of them, you’d have different calculations. Each calculation is a constant at that moment in that specific dataset, but as the dataset process is random, the dataset itself is random, and therefore the calculations are random variables too. There exist as many possible datasets as there are combinations of drawing fixed n units from the fixed N “population” which for both large n and large N is massive. But under central limit theorems, things settle down at some know rates.
All calculations based on the sample are therefore random variables. Regression coefficients are random variables. Standard errors are random variables. T-statistics are random variables. Anything which is a number you calculated based on that specific dataset is paradoxically a random variable under iid sampling. And so we can use that source of randomness to make deductions about the sampling distribution of the estimator across all of the hypothetical samples.
This is the cool part about inference. The t-statistic for instance tells you about the distance between your standard error scaled coefficient and zero. The p-value tells you the share of probability mass that a t-statistic in your sample would appear in a given distribution ordinarily. And so on and so forth, but it’s quite magical and amazes me this was worked out so carefully by so many people going back centuries.
These methods are interesting bc under iid sampling based inference, you’re able to make some meaningful statements about your estimates proximity to the population parameter you care about.
Three kinds of uncertainty: treatment assignment
The second way to quantify uncertainty in our estimates is design-based inference, which Fisher and others developed and which has become increasingly central in modern causal work. Here you hold the sample fixed entirely and ask: what would have happened under a different treatment assignment? The randomization is the source of uncertainty, not the sampling process.
The famous story by Fisher of the lady tasting tea appears to be the origin but perhaps it’s even older. But this approach is the foundation of randomization inference where you perturb, not the sampling process (eg bootstrap, jackknife), but rather you work through the combinations of all possible treatment assignments that could’ve happened, assume a sharp null treatment effect of zero (or some other constant), and then plot the percent of all estimates under alternative assignments that you could’ve gotten. And as before with the t-statistic distribution, yielding its own p-value, here we get the exact p-value that chance would’ve given a measured test statistic as large as the one we got in the real treatment assignment. You have if nowhere else seen this in synthetic control spaghetti plots.
And then there is work by Abadie, Athey, Imbens and Wooldridge that combine both.
Three kinds of uncertainty: researcher/analyst
Both frameworks are elegant and super interesting. I’m teaching probability and statistics this semester and so I am in particular enamored with the sampling approach. It’s really deep, and it’s the source of so many innovations in statistics like the central limit theorem, law of large numbers, bias and consistency.
But the thing about them which I never noticed until I read the many-analyst design papers is that in both sampling and design based inference, the researcher is held fixed. In sampling, you hold fixed n, and you sample it repeatedly in principle which affords us a chance to talk about estimators and estimable in precise ways. In design, you hold the sample fixed, but you work through the reassignment. Both of these allow for precise statements about uncertainty.
But in both of them, you’re still holding fixed the researcher. Neither the standard errors nor the ensuing calculations based on it like t or CI or p-values ever imagine what would’ve happened had someone else worked on the same project as you. Both methodologies treat the researcher as fixed. Neither one is designed to capture what happens when you vary the researcher.
The many-analyst design does. But it isn’t clear to me just what it is that we can pull from it except that there is indeed sources of uncertainty tracing through our sample and assignment to the estimates that comes from the researcher. And not because of publication bias, but rather because of the myriad of decisions that must be made under uncertainty throughout the creation of the analytical sample and the estimates done to it.
When Silberzahn and colleagues sent the same dataset to 29 independent research teams and asked them all the same question, they documented something the profession had been reluctant to quantify: the researcher is not a transparent pipe. The same data, same question, produced a spread of estimates. Not from sampling variation. Not from treatment randomization. But rather from the choices analysts make — which are at least partially endogenous to who they are, what software they trained on, what their advisor told them, what they read last month.
Listen to what they said too at the end about that study:
“These findings suggest that significant variation in the results of analyses of complex data may be difficult to avoid, even by experts with honest intentions. Crowdsourcing data analysis, a strategy in which numerous research teams are recruited to simultaneously investigate the same research question, makes transparent how defensible, yet subjective, analytic choices influence research results.”
See that variation is real. It is literally a source of uncertainty — different team, same data, same question, same experiment, different calculation, different results. Different facts? Different truth? Which is it? Is the answer A or is it B?
We are used to this in some ways. Five people study the minimum wage and come to five conclusions why? Some used city level employment data, some used state panel data, some looked at the UK, some were focused on the 19th century. These were different samples, different treatment assignments, and therefore need not lead to the same estimand.
But when ten researchers working on the same question using the same dataset and the same treatment assignment come to different conclusions, it cannot be any of those things. And the standard errors are correct in the sense that they assume the same team would’ve used every hypothetical sample, but those standard errors don’t measure this other source of uncertainty. But the variation in estimates that hypothetically come from perturbing does.
Now here’s the thing about the many-analyst design as a program: it’s mostly theoretical. You cannot actually send your dataset to 185 independent teams every time you want to publish a paper. When it has been done in these papers, my sense is that it has been to document sources of bias in science. The goal was to document a fact about the world — to prove this third kind of uncertainty exists — not to propose a workflow any individual researcher could follow.
But now I’m wondering otherwise.
What Claude Code changes
The combinatorics of empirical research are staggering in a way that’s easy to underappreciate. Think about it concretely. From raw data cleaning through estimation through table construction, you might face ten major decision points. At each one, you might have two reasonable options. That’s 2^10 possible situations that could’ve occurred by the time the estimates were calculated or 1,024.
If there were 3 options for each of those ten tasks (eg cleaning, measurement), then it’s 3^10 possible ensuing situations or 59,049. That’s 59k different hypothetical estimates.
None of that is usually reported in a study. Researchers historically didn’t even share their code. They did not clearly articulate their design choices. They did not show the robustness of their estimates to dropping this or that differently, including this or that differently. Most do not even probably remember the forks in the road they took to get here. And yet if reasonably different choices could’ve been made or would’ve been made, by a different team, and the calculations at the end would’ve changed as a result, then the estimates are random variables for a third reason than sampling or treatment assignment.
Well here’s the thing. Finding those sources of possible variation is hard automatically. It can be hard for the researcher to see as it’s their own footprints and they may be too close to it. But then working through all those perturbations or a large N randomized sample of them will also be hard. There’s no standardized package to do that, and it’s not clear why we do it, except that here I am noting it is a source of uncertainty, and therefore it’s not clear why it wouldn’t be prioritized when constructing intervals.
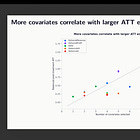
What Claude Code lets you do is automate the perturbation of a subset of those forks. Not all of them — enumerating every discretionary node in a study is itself an unsolved problem, and I’ll come back to that. But you can pick a node. Covariate selection is a natural one, because it is genuinely discretionary — there is no algorithm that tells you which covariates to include to satisfy parallel trends in diff in diff as the parallel trends is not testable. And so different reasonable analysts will make different reasonable choices. Package selection is another, for reasons my experiment made pretty vivid as 75% of the total variation in the outcome came from whether you used python, stata or R (concerning).
The harder problem
There’s a question I raised in the draft of this post and then kind of skated past, so let me come back to it honestly.
Identifying the discretionary nodes — all of them, not just covariate selection — is independently a hard problem. In my experiment I specified the node in advance. I said: the one thing that varies is covariates. That’s a controlled perturbation of one dimension. But in a real study, the discretionary nodes are everywhere and you often don’t know you’re at one. You think you’re making the obvious choice and you don’t realize there were ten other reasonable choices you could have made. The many-analyst design forces you to see that by varying the team.
But I haven’t solved the problem of having Claude enumerate all the discretionary nodes in a given study automatically. That’s something I want to work on, and I think it’s tractable — something like: go through this pipeline step by step and flag every point where a reasonable analyst might have done something different — but I haven’t built that yet.
But I am thinking if you were to construct intervals from analyst uncertainty, then it would be based on the perturbations around endogenous not exogenous nodes. Endogenous nodes are the ones that different researchers would’ve chosen or could’ve chosen. Exogenous nodes do not vary across teams. So coding “African American” as 2 on the race variable might see like a discretionary node but it only would be if there was disagreement. And probably there wouldn’t be for that.
But Claude Code could theoretically find all those nodes. It could find in a pipeline all of the discretionary nodes for you, write the code in a way that perturbs around it and then finds a finite number of “situations” at the end from which estimates are calculated each time. And I think you should be able to work out a p-value. How often is that node pivotal? How often do you find calculations as large as yours?
I think Claude Code could do this for us, do it fast, and correctly. I think we could report a forest plot of these estimates.
What I am not sure of if the large sample properties of large N teams. It’s not clear to me why it wouldn’t follow the same central limit theorems as the rest but I guess I pause because it’s entirely clear what an estimand is if there is alternative measurement / package choices one could make in a given sample.



I have three thoughts, as someone who is not reading the Claude Code entries but is interested in the broader issue to you bring up here.
1) How on earth is the software generating 75% of the variation? You put (concerning) where I would put (terrifying!!!). It's hard to imagine variation in tiebreaking rules and such would be enough to do that.
2) I think you should give empirical researchers more credit. While we haven't integrated this source of variation into our standard errors, it is completely normal (and expected) to revisit as many forks in the road as possible and see if results change appreciably. Current papers often have a visualization with 30+ alternative specifications all together on one graph showing how much this variation in reasonable specification or measurement choices seems to matter. It is not formally integrated into standard errors, but it is there to help us see whether we should be more or less confident that we've got reasonable bounds on betahat. Getting the inference right is more important to me than being sure my final, singular p-value from the "main estimate" is corrected for this kind of error.
3) Which brings me to the last thing. I am not sure I want to think of these as error in the same sense as the other two sources. Some set (maybe a large set) of the different specifications are biased because they do not meet the (untestable) assumptions about selection on observables or whatever. I'm sort of reminded of the conversation between Lalonde 86 and Heckman and ?? (maybe Robb?). Lalonde says "oh my goodness, the vast majority of non-experimental estimates are garbage" and Heckman says "well, sure, but we could have thrown out half of them ex-ante as garbage, so the success rate isn't as bad as you think." I mean, letting the program select covariates means you are certainly comparing poorly-specified to better-specified options. It may be hard to know which are which, but my point is: it's not a natural source of variation but it's what happens when you mix unbiased and biased estimates together and the look at the distribution. I'm mean, perhaps it's error in the most real sense, that some of these estimates are erroneous!
I may be missing the boat with this last one, if you have reason to think that all of the coding processes are ruling out all the biased estimates, but I guess I'm not convinced they can be distinguished. Wheat and weeds growing up entangled together in the field, right? So I'm not saying we shouldn't care about this error, just that I don't see it as having the same sort of statistical properties that would allow us to work it into our standard errors somehow. Or maybe I'm saying, I think looking at those crazy robustness figures isn't a bad substitute for some formalization of this that would treat different estimates symmetrically.
Simonsohn/Simmons/Nelson suggest a combined bootstrap-specification inference procedure — is that on the lines you’re thinking of? http://urisohn.com/sohn_files/wp/wordpress/wp-content/uploads/specification-curve-published-hand-corrected.pdf